Food & Drink
San Diego's Best Restaurants of 2024
Read article
Featured articles

Food News
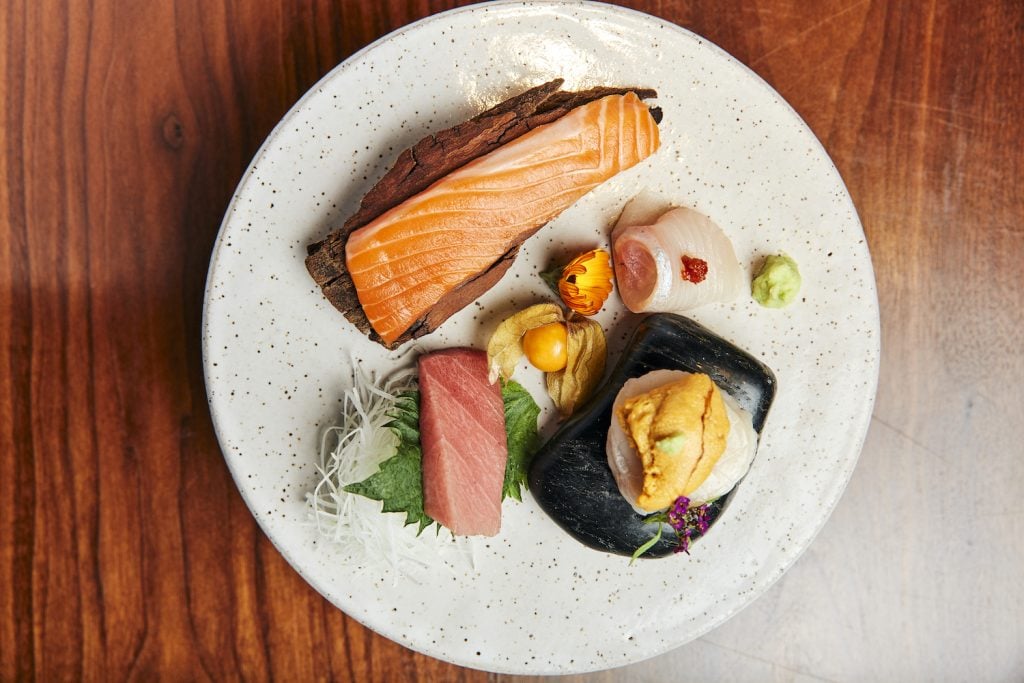

Food & Drink
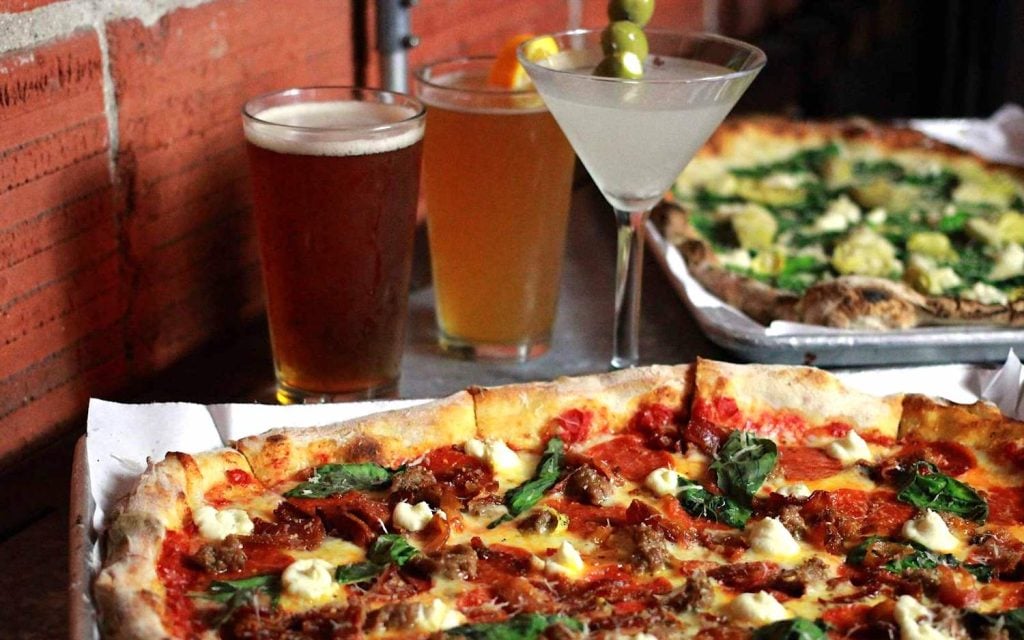

Food & Drink
Featured articles
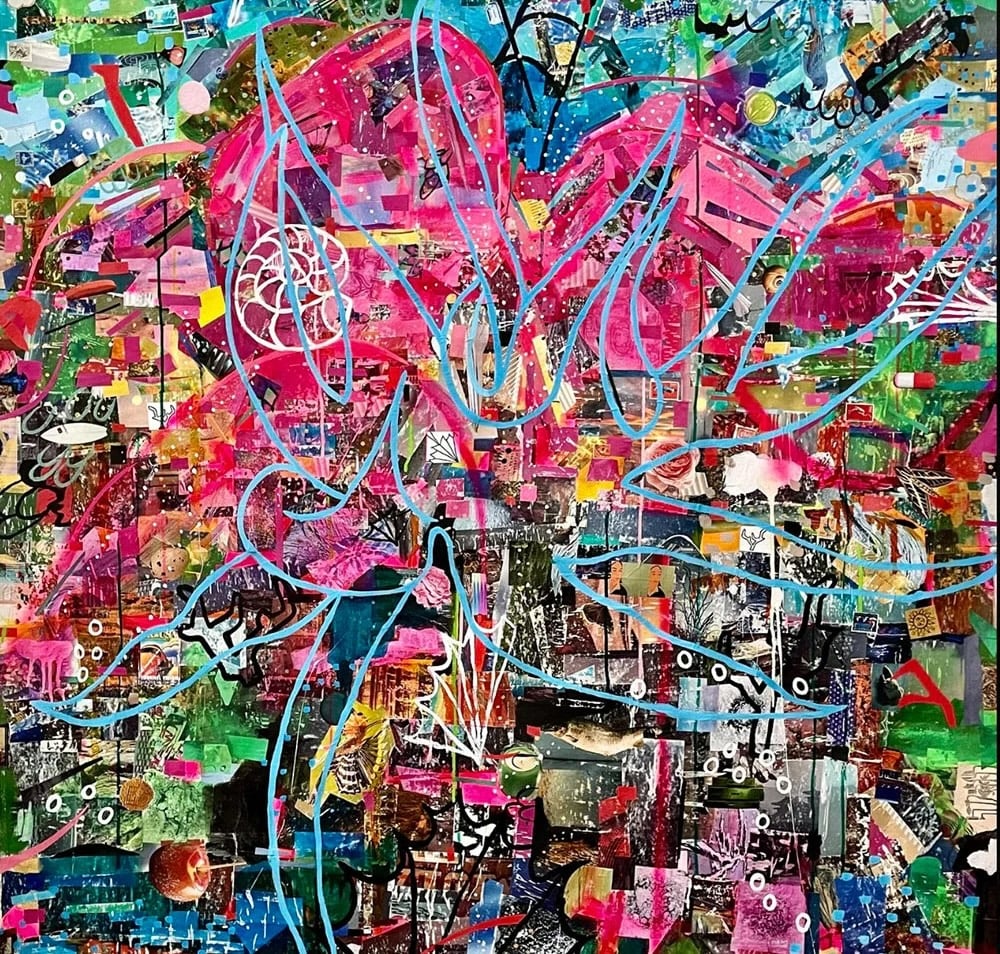

Arts & Culture

Food & Drink
Everything SD
Featured articles

Arts & Culture
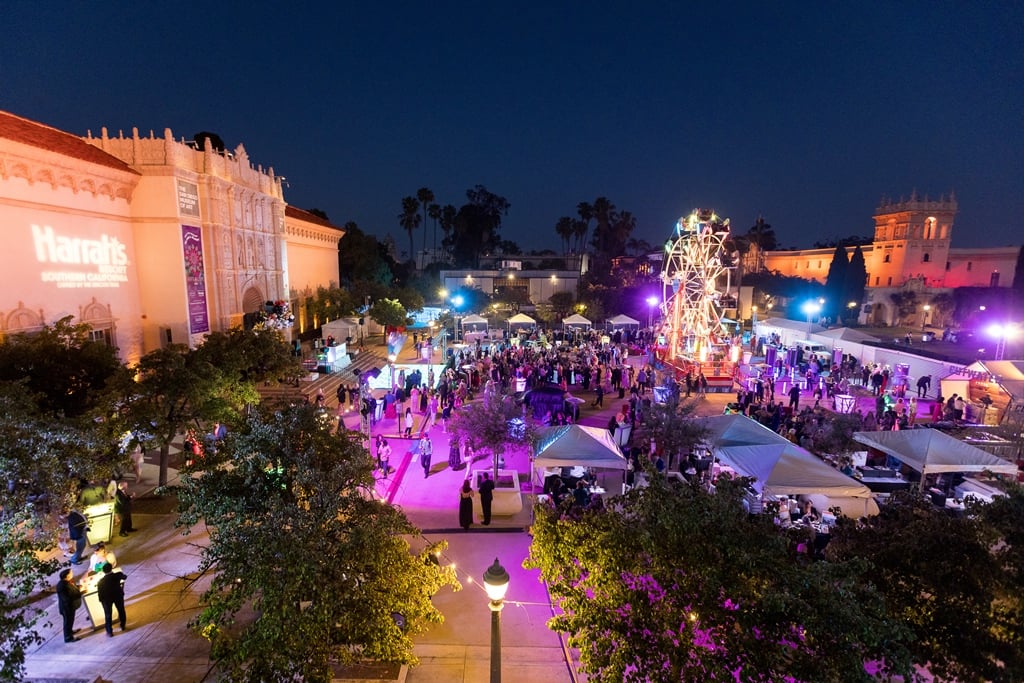

Things to Do
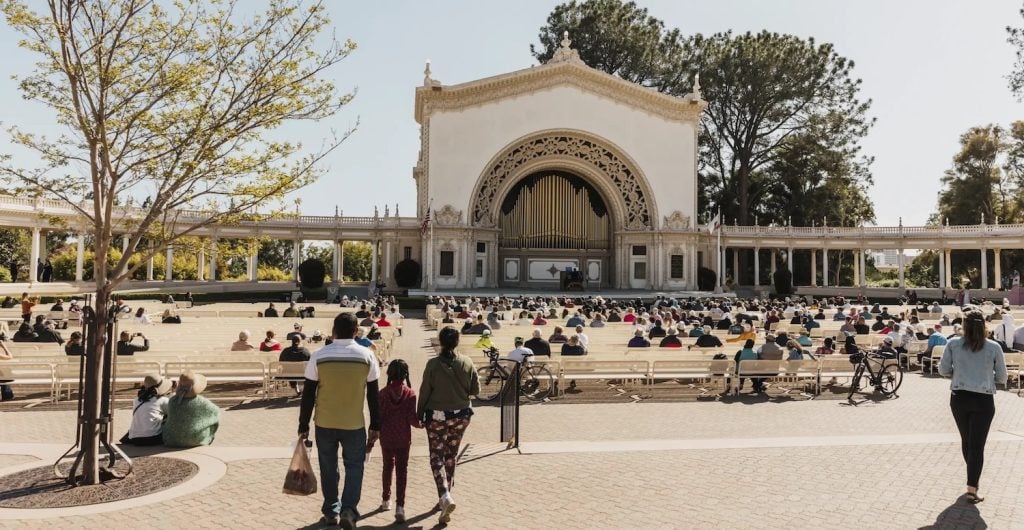

Things to Do
Featured articles

podcast-ep

podcast-ep

podcast-ep
Featured articles

Arts & Culture

Food & Drink

Features
Ready to know more about San Diego?
SubscribeReady to know more about San Diego?
Food & Drink
The longtime joint on Tenth Ave. will exit its location after baseball season with plans to reopen at Park 12 development
Read article
Food & Drink
The movers and shakers revolutionizing our city's restaurants and bars
Read article
Food & Drink
The chef and owner of the popular Golden Hill taco truck is setting down roots in the former Krakatoa space this summer
Read article
Food & Drink
Plus how local craft beer measures up nationally, upcoming chef dinners with Drew Deckman, and more food and drink news
Read article
Everything SD
Editor Nicolle Monico shares five insights she’s gained over the last few weeks while dating in San Diego
Read article
Everything SD
The San Diego curator’s paintings are on view at Tijuana’s Sala de Espera gallery through May 4
Read article
Everything SD
With a homegrown talent program, the newest team is town appears to be building to win
Read article
Everything SD
Is financial stability just another excuse to stay guarded or a valid concern while dating?
Read article
Features
How a quiet kid from Oceanside went from sleeping in a parking lot to becoming one of the nation's biggest personalities
Read article
Features
The local couple collecting rare and bygone buds, including one of the most famous urban legends in cannabis culture
Read article
Features
Designer Susan Wintersteen of Savvy Interiors walks us through her favorite kitchen projects
Read article
Features
Spruce up your surrounds (and yourself) with our curated list of San Diego products
Read article
Arts & Culture
How to stay busy and important this month in America's Finest City
Read article
Things to Do
Ride on down to the rodeo in Lakeside, admire art-inspired floral arrangements at Art Alive, and vote for the city’s best cocktail in the Gaslamp
Read article
Things to Do
Escape the house for some family fun without busting your budget
Read article
Things to Do
Vote for the city’s best salsa in North Park, celebrate classic cars in style in La Jolla, and keep San Diego green at the Creek to Bay Cleanup
Read article
PODCAST
After Covid crushed his baseball dreams, a kid from Oceanside applied his relentless work ethic to amassing tens of millions of food-focused followers
Read article
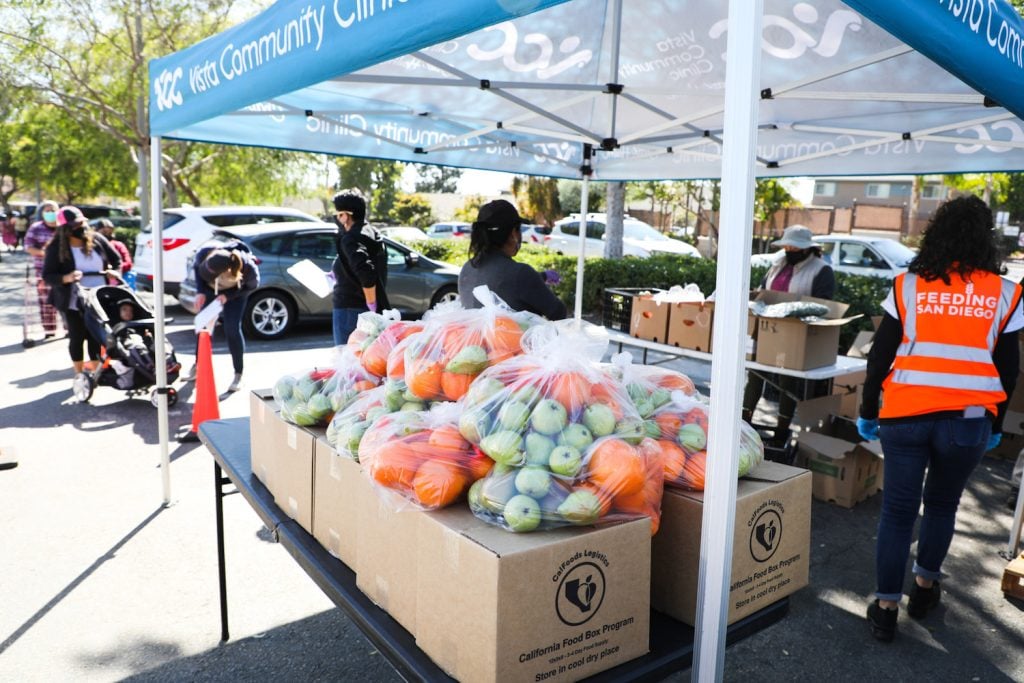
PODCAST
The city’s leading nonprofit tackling hunger joins us on the podcast this week
Read article
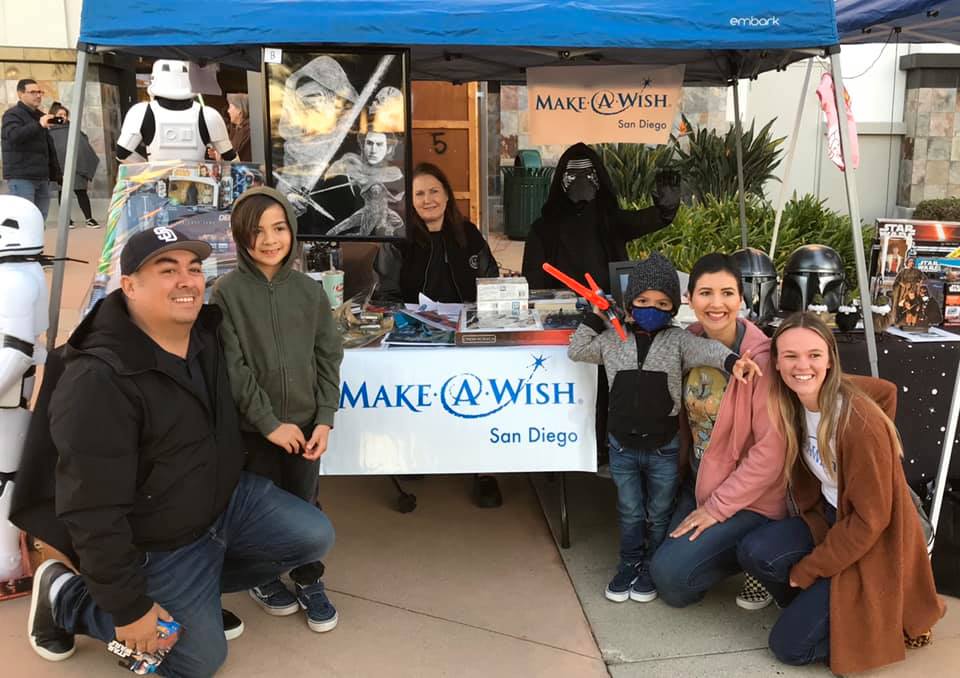
PODCAST
Make-A-Wish joins the podcast to discuss its new partnership with Matsu's executive chef/owner & upcoming gala
Read article
video
The social media sugar sensation opened a new flagship location with much more than donuts.
Read article

Partner content
Read article

Partner content
Read article

Partner content
Read article

Partner content
Read article
Subscribe to our Newsletters
By clicking Subscribe you’re confirming that you agree with our Terms and Conditions.
Email: [email protected]

© Copyright 2023 San Diego Magazine 1230 Columbia Street, Suite 800, San Diego, CA